neuror cut-plane¶
CLI utilities related to cut-plane repair.
neuror cut-plane [OPTIONS] COMMAND [ARGS]...
compute¶
CLI utilities to compute cut planes.
neuror cut-plane compute [OPTIONS] COMMAND [ARGS]...
file¶
Find the position of the cut plane (it assumes the plane is aligned along X, Y or Z) for morphology FILENAME.
It returns the cut plane and the positions of all cut terminations.
Compute a cut plane and outputs it either as a STDOUT stream or in a file
if -o option is passed.
The control plots can be displayed by passing the -d option.
The bin width can be changed with the -w option (see below)
Description of the algorithm:
The distribution of all points along X, Y and Z is computed and put into 3 histograms.
For each histogram we look at the first and last empty bins (that is, the last bin before the histogram starts rising, and the first after it reaches zero again). Under the assumption that there is no cut plane, the posteriori probability of observing this empty bin given the value of the non-empty neighbour bin is then computed.
The lowest probability of the 6 probabilities (2 for each axes) corresponds to the cut plane.
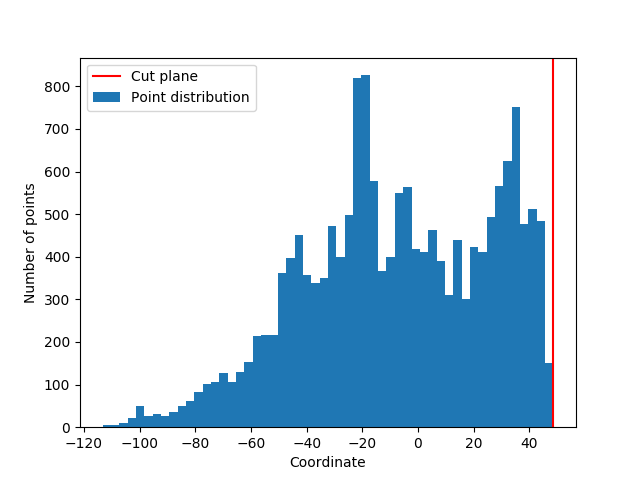
- Returns:
A dictionary with the following items:
- status:
‘ok’ if everything went right, else an informative string
- cut_plane:
a tuple (plane, position) where ‘plane’ is ‘X’, ‘Y’ or ‘Z’ and ‘position’ is the position
- cut_leaves:
an np.array of all termination points in the cut plane
- figures:
if ‘display’ option was used, a dict where values are tuples (fig, ax) for each figure
- details:
A dict currently only containing -LogP of the bin where the cut plane was found
Example:
neuror cut-plane compute file -d tests/data/Neuron_slice.h5 -o my-plane.json -w 10
neuror cut-plane compute file [OPTIONS] FILENAME
Options
- -o, --output <output>¶
Output name for the JSON file (default=STDOUT)
- -w, --width <width>¶
The bin width (in um) of the 1D distributions
- -d, --display¶
Flag to enable the display control plots
- -p, --plane <plane>¶
Force the detection along the given plane
- Options:
x | y | z
- --position <position>¶
Force the position. Requires –plane to be set as well
Arguments
- FILENAME¶
Required argument
folder¶
Compute cut planes for all morphology in INPUT_DIR and save them into OUTPUT_DIR
See “cut-plane compute –help” for more information
neuror cut-plane compute folder [OPTIONS] INPUT_DIR OUTPUT_DIR
Options
- -w, --width <width>¶
The bin width (in um) of the 1D distributions
- -d, --display¶
Flag to enable the display control plots
- -p, --plane <plane>¶
Force the detection along the given plane
- Options:
x | y | z
Arguments
- INPUT_DIR¶
Required argument
- OUTPUT_DIR¶
Required argument
hint¶
Launch the app to manually search for the cut plane. After running the command, either click the link in the console or open your browser and go to the address shown in the console.
Example:
neuror cut-plane compute hint ./tests/data/Neuron_slice.h5
neuror cut-plane compute hint [OPTIONS] FILENAME
Arguments
- FILENAME¶
Required argument
join¶
Merge cut-planes from json files located at PLANE_PATHS into one. The output is writen at OUT_FILENAME
Example:
neuror cut-plane join result.json plane1.json plane2.json plane3.json
neuror cut-plane join [OPTIONS] OUT_FILENAME [PLANE_PATHS]...
Arguments
- OUT_FILENAME¶
Required argument
- PLANE_PATHS¶
Optional argument(s)
repair¶
CLI utilities to repair cut planes.
neuror cut-plane repair [OPTIONS] COMMAND [ARGS]...
file¶
Repair dendrites of a cut neuron.
neuror cut-plane repair file [OPTIONS] INPUT_FILE OUTPUT_FILE
Options
- --plot_file <plot_file>¶
Where to save the plot
- -a, --axon-donor <axon_donor>¶
A morphology that provides a reference axon
- --cut-file <cut_file>¶
Path to a CSV whose columns represents the X, Y and Z coordinates of points from which to start the repair
Arguments
- INPUT_FILE¶
Required argument
- OUTPUT_FILE¶
Required argument
folder¶
Repair dendrites of all neurons in a directory.
neuror cut-plane repair folder [OPTIONS] INPUT_DIR OUTPUT_DIR
Options
- --plot_dir <plot_dir>¶
- -a, --axon-donor <axon_donor>¶
A morphology that provides a reference axon
- --cut-file-dir <cut_file_dir>¶
A dir with the cut points CSV file for each morphology. See also “neuror cut-plane repair file –help”.
Arguments
- INPUT_DIR¶
Required argument
- OUTPUT_DIR¶
Required argument